3种机器学习方法同时结合预后生存期展开分析!确定不进来看看?(二)
点击蓝字 关注我们
小伙伴们大家好啊,今天大海哥将要继续接着上次没有结束的分享接着说,3种机器学习算法同时结合预后生存期分析!很多小伙伴肯定等不及了,快开始吧!等了好几天了都,先别着急,为了怕很多小伙伴忘记上一次咱们到底做了什么,大海哥就先介绍一下上次咱们主要的分析结果,然后再来深入学习今天的内容~让我们开始吧!
上一次呢?我们通过数据预处理以及数据清洗剔除掉了一些冗余的字段,同时计算了每个变量缺失数据的所占百分比,然后删除缺失值较多的列,同时把一些关键变量转为因子并且删除只具有一个属性的因子数据。数据处理完后,我们还通过探索性分析来查看了变量之间的相关性以及一些连续变量伴随生存结果的箱型图,最后针对肿瘤尺寸和突变的数据存在很多极大值的情况,对这两个字段进行了log10处理,使得数据整体符合正态分布。
好了,上述就是上一次我们所做的事情,那么今天我们就要拿着处理好的数据,来进行分析啦!首先同样的导入我们所需要的R包。
#加载R包library(corrplot)library(VIM)library(ggplot2)library(gridExtra)library(scales)library(tidyr)library(dplyr)library(readr)library(ggformula)library(imputeMissings)library(gapminder)library(caret)library(rpart)library(rattle)library(kernlab)library(randomForest)library(DMwR2)library(car)library(MLeval)#导入数据#然后就是加载上一次分析得到的处理后的数据clinical <- read.csv("clinical.csv", header=TRUE, sep=",", stringsAsFactors = FALSE)clinical_e<- clinicaldim(clinical_e)
经过预处理后,我们还有20个特征,看看都是些什么

#然后我们需要重新定义一下木匾变量,就是overall_survival列make.names("overall_survival", unique = TRUE, allow_ = FALSE);clinical_e %>%mutate(overall_survival = factor(overall_survival,labels = make.names(levels(overall_survival))));levels(clinical_e$overall_survival) <- c("D", "L")#死亡就是D,存活就是L#本次分析呢,就是通过随机森林和支持向量机 (SVM) 预测癌症患者是否会生存,其中SVM包含两种核函数即多项式核和径向核。#首先我们要在随机森林和SVM上进行单层10倍交叉验证。选择每次分割 (mtry) 时随机采样作为候选变量的数量来优化随机森林模型。“degree of the polynomial”和cost分别在多项式和径向核中分别进行调整。#话不多说,创建模型吧!set.seed(789)#设置随机种子#定义交叉验证参数ctrl = trainControl(method = "cv", number = 10,savePredictions = TRUE,classProbs = TRUE,verboseIter = TRUE)#定义一个非分类的交叉验证参数ctrl_1 = trainControl(method = "cv", number = 10)#保留一下数据data_used = clinical_edata_used_1 = clinical#设置多项式核SVM交叉验证fit_poly = train(overall_survival ~ .,data = data_used,na.action = na.exclude,method = "svmPoly",tuneGrid = expand.grid(C = c(1),degree = seq(0, 5, by = 1), scale = c(1)), # gammapreProcess = c("center","scale"),trControl = ctrl)#看一下交叉验证结果,给出了最优参数,就在图片最后一行fit_poly

#设置径向核SVM交叉验证fit_radial = train(overall_survival ~ .,data = data_used,method = "svmRadial",tuneGrid = expand.grid(C = c(1:5), sigma = c(1)),preProcess = c("center","scale"),trControl = ctrl)fit_radial

#设置随机森林交叉验证fit_rf = train(overall_survival ~.,data = data_used,method = "rf",na.action = na.exclude,tuneGrid = expand.grid(mtry = c(3:12)),preProcess = c("center","scale"),trControl = ctrl)fit_rf
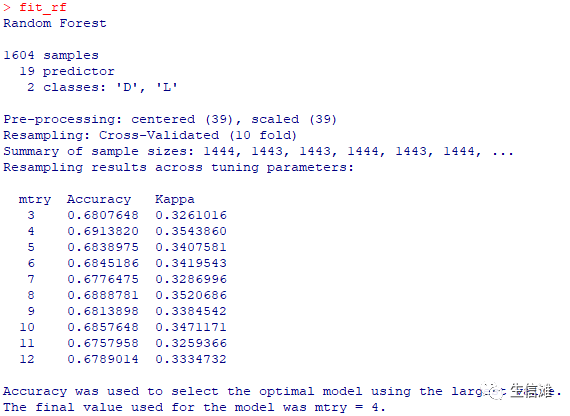
#定义不分类随机森林交叉验证,这个结果没有上面那个好,那就选上面那个fit_rf_1 = train(overall_survival ~.,data = data_used_1,method = "rf",na.action = na.exclude,tuneGrid = expand.grid(mtry = c(3:12)),preProcess = c("center","scale"),trControl = ctrl_1)
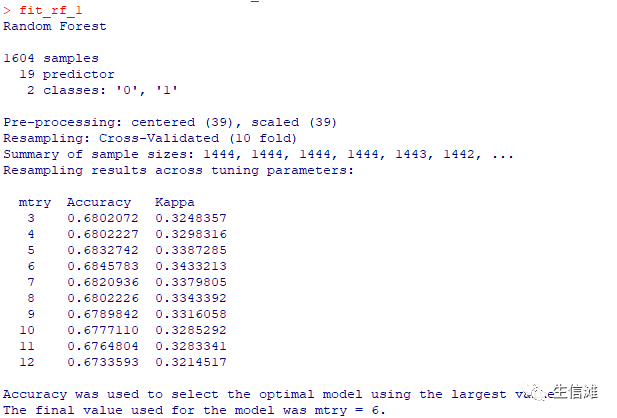
all_best_Types = c("Polynomial Kernel","Radial Kernel","Random forests")all_best_Pars = list(fit_poly$bestTune, fit_radial$bestTune,fit_rf$bestTune )all_best_Models = list(fit_poly,fit_radial,fit_rf)all_best_Accuracy = c(max(fit_poly$results$Accuracy),max(fit_radial$results$Accuracy),max(fit_rf$results$Accuracy))############# 模型对比############## 看看特征数和准确率的关系mtree = length(seq(0, 5, by = 1));mtree1 = length(seq(1, 5, by = 1))mtree2 = length(seq(3, 12, by = 1))mmodels = mtree+mtree1+mtree2modelMethod = c(rep("Polynomial Kernel",mtree),rep("Radial Kernel",mtree1), rep("Random forests",mtree2))all_caret_Accuracy = c((fit_poly$results$Accuracy),fit_radial$results$Accuracy, fit_rf$results$Accuracy)coloptions = rainbow(4)colused = coloptions[as.numeric(factor(modelMethod))]charused = 5*(as.numeric(factor(modelMethod)))#可视化结果看看plot(1:mmodels,all_caret_Accuracy,col=colused,pch=charused,xlab = "Model label",ylab = "Accuracy")order.min = c(which.max(fit_poly$results$Accuracy),mtree+which.max(fit_radial$results$Accuracy),mtree+mtree-1+which.max(fit_rf$results$Accuracy))abline(v=order.min,lwd=0.5)abline(v=order.min[2],col="red",lwd=0.5)abline(v=order.min[3],col="green",lwd=0.5)
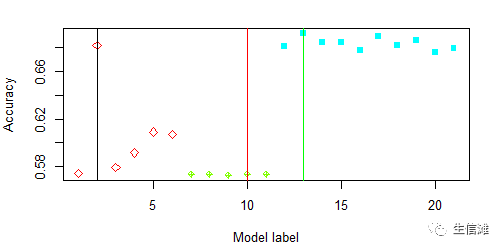
#整体看的话,还是随机森林要更好一点,多项式核SVM不太稳定,径向核SVM稳定,但是ACC太低了#然后就要画一下ROC曲线了,毕竟这是大家最常用的评估手段了#这里会产生很多图,可以获取每个模型的关键信息,这里大海哥就不展示了poly <- evalm(fit_poly);radial <- evalm(fit_radial);rf <- evalm(fit_rf);#获取我们想要的信息fpr = poly$roc$data$FPRpo = poly$roc$data$SENSra = radial$roc$data$SENSrff = rf$roc$data$SENSroc_fig <- data_frame(fpr,po,ra, rff)require(ggplot2)colors <- c("Polynomial Kernel" = "blue", "Radial Kernel" = "red", "Random forests" = "green")ggplot(roc_fig, aes(fpr)) +geom_line(aes(y=po, colour="Polynomial Kernel"), size = 1) +geom_line(aes(y=ra, colour="Radial Kernel"), size = 1)+geom_line(aes(y=rff, colour="Random forests"), size = 1)+geom_abline(intercept = 0, slope = 1, col = "gray")+labs(title ="ROC curve",x = "Specificities",y = "Sensitivities",color = "Legend") +scale_color_manual(values = colors)
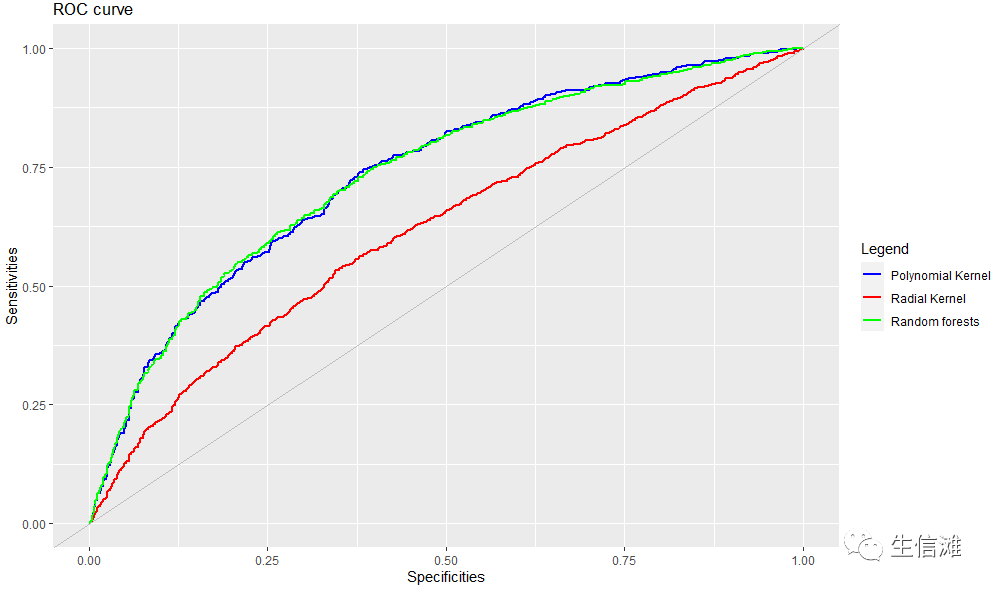
#这里看ROC,多项式核SVM和随机森林不相伯仲!Model = c("Polynomial Kernel","Radial Kernel","Random forests")AUC_Val = c(poly$stdres$`Group 1`$Score[13],radial$stdres$`Group 1`$Score[13], rf$stdres$`Group 1`$Score[13])#同时我们还整合了ACC值,一起看一下df2 <- data.frame(supp=rep(c("Accuracy", "AUC"), each=3),mod=rep(Model,2),va=c(all_best_Accuracy, AUC_Val))ggplot(data=df2, aes(x=mod, y=va, fill=supp)) +geom_bar(stat="identity", position=position_dodge())+labs(x = "",y = "Value",fill = "")
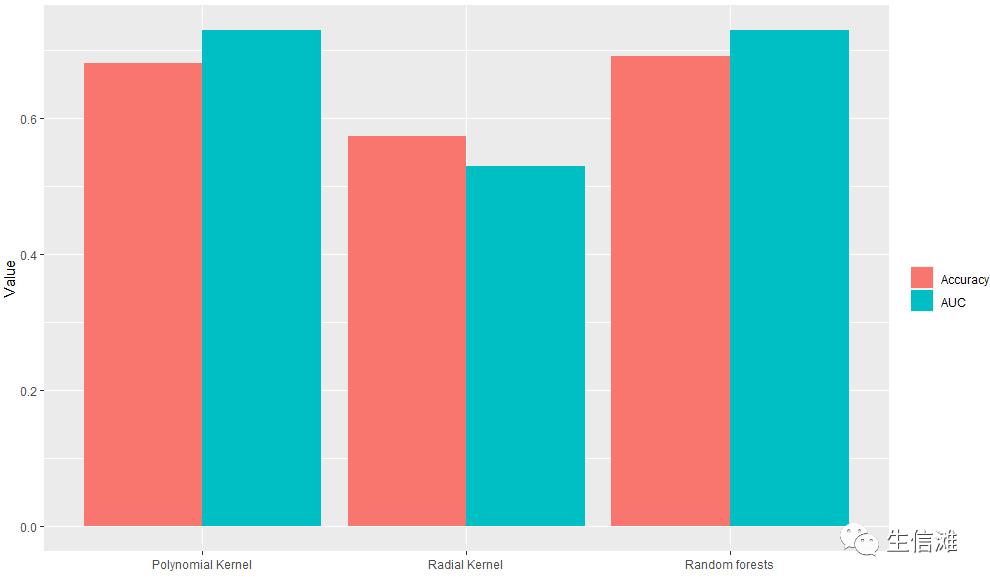
#但是ACC好像随机森林更优!#看看随机森林的重要性评分importance(fit_rf_1$finalModel)varImpPlot(fit_rf$finalModel)
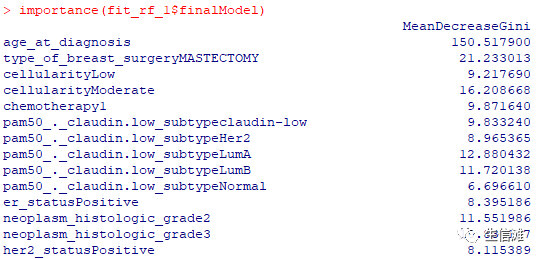
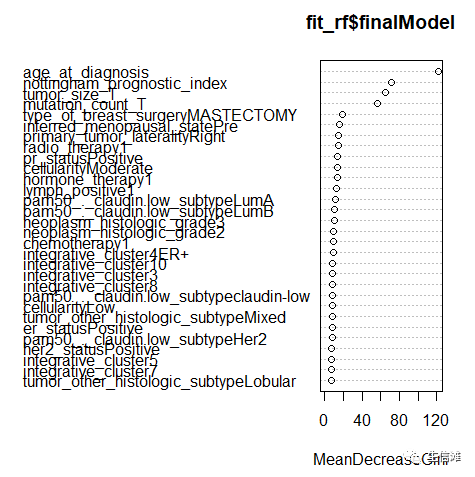
因为我们处理过年龄字段,所以可以得出结论,如果患者诊断时的年龄小于50岁,她的生存概率很高。对于 50 岁以上的患者,我们看到生存机会发生急剧变化。如果患者的年龄超过75岁,她的生存机会非常低,并且死亡的可能性很大。
好了,这次的分析到这里就结束啦!如此完整的一套多机器学习预测预后的分析,你确定不自己尝试一下?动手试试吧!
(最后推荐一下大海哥新开发的零代码云生信分析工具平台包含超多零代码小工具,上传数据一键出图,感兴趣的小伙伴欢迎来参观哟。
网址:
http://www.biocloudservice.com/home.html)

生信滩公众号

点击“阅读原文”进入网址
